---
There can be no doubt that the term species denotes a set composed of organisms which can interbreed with one another, thus perpetuating an evolving spatiotemporal principle to a new generation, of which is composed a new set of unique individuals, operating and maintaining novel processes in their life-history. Largely, this firmness of belief stems from the operational usages of the term “species” in evolutionary biology. The existence of algorithms, such as hierarchical clustering, which can split populations of individuals into taxa, implies a real, although abstract, way of distinguishing finite attributes across a list of objects. Generally, hierarchical clustering, also called numerical taxonomy, is the process of producing groups of objects which share statistically meaningful pair-wise relationships among a list of chosen attributes for each object (2). The taxa that are generated by such a process can be as multi-dimensional as one would like, although the clustering of individuals into apparent classes should occur regardless of the system you are focusing on. As an example, clustering by genes would produce the same splitting as a clustering by phenotypic characteristics such as “wing length” and “head circumference” as measured in a group of some birds, although an observer could actually take both systems (the genetic and phenotypic) into account as variables assigned to their objects of study. I reject the notion that the entire set of such individuals is actually a natural kind in the sense that gold is a natural kind, as this multi-dimensional space in biological life is constantly evolving through time. Indeed, if one were to collect information about individual objects composing the class of all objects falling under “gold”, you may find over time that such a list of characteristics are also changing. The difference, perhaps, is that the method of clustering you choose to classify objects such as gold versus biological life produces a significant difference in variance. The change over thousands versus millions of years in gold apparently does not exhibit the novelty that biological life does at those time-scales.
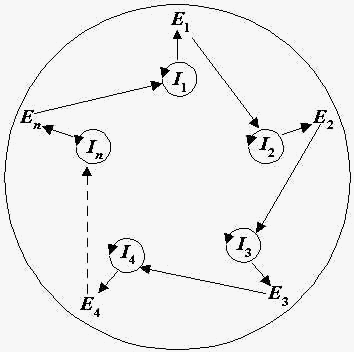
Certainly, when you recognize that this method of clustering can take into consideration many levels of resolution, it must be recognized that the splitting of a feature-space into taxa will occur at many levels of resolution as well. Where do we distinguish applying the word species merely to a study involving the clustering of cells within an individual into groups as opposed to comparing actual biological individuals against one another? Once again, I would pretend to claim that this becomes operational matter. If it is experimentally useful to analyze the internal machinations of an individual and then assign observable clusters produced by them the name of “species”, then the observer is certainly entitled to do so. However, this must be distinguished from the act of how we consider a species in the domain of evolutionary biology; without that distinction, we would have no basis for considering when Homo sapiens diverged from its ancestral forms. Indeed, this notion we recognize as speciation is a benchmark of evolutionary theory. The other alternative is unifying these multiple resolutions by doing the following. If a biological individual is composed of cells, and natural selection posits that the fittest forms of life pass their spatiotemporal principle to a new generation, then the taxa of cells which continue to proliferate from a cluster x are the fittest for that individual. This cluster will continue to proliferate until such a time when the individual dies. Down to the resolution of the individual cell, the makeup that composes that cell from its original point of origin for the ontogeny of that organism from birth also uniquely proliferates insofar as being the informational element that initiates and survived from the moment of conception. Applying natural selection to the scale of unique components such as nucleic acids and proteins should also show us that only the fittest of them will continue to grow, whereas ones which contribute to those processes running counter to self-renewal would be discarded. We can continue an analysis in this fashion at all scales of resolution and organize the top clusters which have survived into the present into the nodes of what is called Eigen’s hypercycle (1). Such a hypercycle is organizationally closed, which means that all components are involved in a self-renewing process working towards maintaining the existence of the individual. The interaction also creates a closure between the individuals of the set called “species” which they are involved in, and this interaction is the ability of interbreeding at the level of resolution for a group of organisms. The hypercycle which survives is that closure between individuals in a set that manage to exist in the future, as well. In evolutionary time, once an observer in the future can note sufficient variance between these unique cycles at time-point X1 and X2, she may also recognize enough noticeable differences, whether from genetic analysis or the fossil record, that speciation has occurred. This type of analysis and recognition of apparently unique classes is something uncharacteristic of a natural kind and rejects the notion of an essential force underlying individuals or groups of them. Another particularly beautiful result of such a method also places the species problem as an ontological one. In the world of biology, ontological solutions in the questions of life have certainly allowed the scientists to at least deal with the deluge of data and biological information generated in the laboratory.
REFERENCES
1. Eigen, M., and P. Schuster. 1978. “Part A: Emergence of the Hypercycle.” Naturwissenschaften 65:7–41.
2. Press, WH; Teukolsky, SA; Vetterling, WT; Flannery, BP (2007). "Section 16.4. Hierarchical Clustering by Phylogenetic Trees". Numerical Recipes: The Art of Scientific Computing (3rd ed.). New York: Cambridge University Press.
There can be no doubt that the term species denotes a set composed of organisms which can interbreed with one another, thus perpetuating an evolving spatiotemporal principle to a new generation, of which is composed a new set of unique individuals, operating and maintaining novel processes in their life-history. Largely, this firmness of belief stems from the operational usages of the term “species” in evolutionary biology. The existence of algorithms, such as hierarchical clustering, which can split populations of individuals into taxa, implies a real, although abstract, way of distinguishing finite attributes across a list of objects. Generally, hierarchical clustering, also called numerical taxonomy, is the process of producing groups of objects which share statistically meaningful pair-wise relationships among a list of chosen attributes for each object (2). The taxa that are generated by such a process can be as multi-dimensional as one would like, although the clustering of individuals into apparent classes should occur regardless of the system you are focusing on. As an example, clustering by genes would produce the same splitting as a clustering by phenotypic characteristics such as “wing length” and “head circumference” as measured in a group of some birds, although an observer could actually take both systems (the genetic and phenotypic) into account as variables assigned to their objects of study. I reject the notion that the entire set of such individuals is actually a natural kind in the sense that gold is a natural kind, as this multi-dimensional space in biological life is constantly evolving through time. Indeed, if one were to collect information about individual objects composing the class of all objects falling under “gold”, you may find over time that such a list of characteristics are also changing. The difference, perhaps, is that the method of clustering you choose to classify objects such as gold versus biological life produces a significant difference in variance. The change over thousands versus millions of years in gold apparently does not exhibit the novelty that biological life does at those time-scales.
Certainly, when you recognize that this method of clustering can take into consideration many levels of resolution, it must be recognized that the splitting of a feature-space into taxa will occur at many levels of resolution as well. Where do we distinguish applying the word species merely to a study involving the clustering of cells within an individual into groups as opposed to comparing actual biological individuals against one another? Once again, I would pretend to claim that this becomes operational matter. If it is experimentally useful to analyze the internal machinations of an individual and then assign observable clusters produced by them the name of “species”, then the observer is certainly entitled to do so. However, this must be distinguished from the act of how we consider a species in the domain of evolutionary biology; without that distinction, we would have no basis for considering when Homo sapiens diverged from its ancestral forms. Indeed, this notion we recognize as speciation is a benchmark of evolutionary theory. The other alternative is unifying these multiple resolutions by doing the following. If a biological individual is composed of cells, and natural selection posits that the fittest forms of life pass their spatiotemporal principle to a new generation, then the taxa of cells which continue to proliferate from a cluster x are the fittest for that individual. This cluster will continue to proliferate until such a time when the individual dies. Down to the resolution of the individual cell, the makeup that composes that cell from its original point of origin for the ontogeny of that organism from birth also uniquely proliferates insofar as being the informational element that initiates and survived from the moment of conception. Applying natural selection to the scale of unique components such as nucleic acids and proteins should also show us that only the fittest of them will continue to grow, whereas ones which contribute to those processes running counter to self-renewal would be discarded. We can continue an analysis in this fashion at all scales of resolution and organize the top clusters which have survived into the present into the nodes of what is called Eigen’s hypercycle (1). Such a hypercycle is organizationally closed, which means that all components are involved in a self-renewing process working towards maintaining the existence of the individual. The interaction also creates a closure between the individuals of the set called “species” which they are involved in, and this interaction is the ability of interbreeding at the level of resolution for a group of organisms. The hypercycle which survives is that closure between individuals in a set that manage to exist in the future, as well. In evolutionary time, once an observer in the future can note sufficient variance between these unique cycles at time-point X1 and X2, she may also recognize enough noticeable differences, whether from genetic analysis or the fossil record, that speciation has occurred. This type of analysis and recognition of apparently unique classes is something uncharacteristic of a natural kind and rejects the notion of an essential force underlying individuals or groups of them. Another particularly beautiful result of such a method also places the species problem as an ontological one. In the world of biology, ontological solutions in the questions of life have certainly allowed the scientists to at least deal with the deluge of data and biological information generated in the laboratory.
REFERENCES
1. Eigen, M., and P. Schuster. 1978. “Part A: Emergence of the Hypercycle.” Naturwissenschaften 65:7–41.
2. Press, WH; Teukolsky, SA; Vetterling, WT; Flannery, BP (2007). "Section 16.4. Hierarchical Clustering by Phylogenetic Trees". Numerical Recipes: The Art of Scientific Computing (3rd ed.). New York: Cambridge University Press.
No comments:
Post a Comment